It’s hardly ever entirely one extreme or the other in natural language generation and controlled natural languages. Rarely can one get away with simplistic ‘just fill in the blanks’ templates that do not do any grammar or phonological processing to make the output better; our technical report about work done some 17 years ago was a case in point on the limitations thereof if one still needs to be convinced [1]. But where does NLG start? I agree with Ehud Reiter that it isn’t about template versus NLG, but a case of levels of sophistication: the fill-in-the-blank templates definitely don’t count as NLG and full-fledged grammar-only systems definitely do, with anything in-between a grey area. Adding word-level grammatical functions to templates makes it lean to NLG, or even indeed being so if there are relatively many such rules, and dynamically creating nicely readable sentences with aggregation and connectives counts as NLG for sure, too.
With that in mind, we struggled with how to name the beasts we had created for generating sentences in isiZulu [2], a Niger-Congo B language: nearly every resultant word in the generated sentences required a number of grammar rules to make it render sufficiently well (i.e., at least grammatically acceptable and understandable). Since we didn’t have a proper grammar engine yet, but we knew they could never be fill-in-the-blank templates either, we dubbed them verbalisation patterns. Most systems (by number of systems) use either only templates or templates+grammar, so our implemented system [3] was in good company. It may sound like oldskool technology, but you go ask Meta with their Galactica if a ML/DL-based approach is great for generating sensible text that doesn’t hallucinate… and does it well for languages other than English.
That said, honestly, those first attempts we did for isiZulu were not ideal for reusability and maintainability – that was not the focus – and it opened up another can of worms: how do you link templates to (partial) grammar rules? With the ‘partial’ motivated by taking it one step at a time in grammar engine development, as a sort of agile engine development process that is relevant especially for languages that are not well-resourced.
We looked into this recently. There turn out to be three key mechanisms for linking templates to computational grammar rules: embedding (E), where grammar rules are mixed with the templates specifications and therewith co-dependent, and compulsory (C) and partial (P) attachment where there is, or can be, an independent existence of the grammar rules.

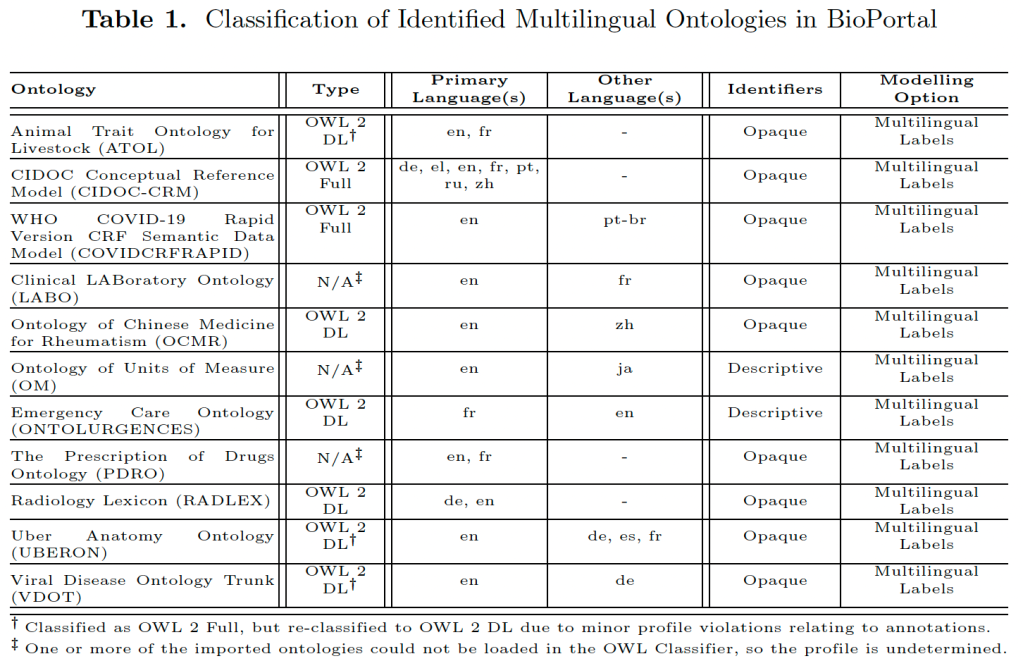
The difference between the latter two is subtle but important for use and reuse of grammar rules in the software system and the NLG-ness of it: if each template must use at least one rule from the set of grammar rules and each rule is used somewhere, then the set of rules is compulsorily attached. Conversely, it is partially attached if there are templates in that system that don’t have any grammar rules attached. Whether it is partial because it’s not needed (e.g., the natural language’s grammar is pretty basic) or because the system is on the fill-in-the-blank not-NLG end of the spectrum, is a separate question, but for sure the compulsory one is more on the NLG side of things. Also, a system may use more than one of them in different places; e.g., EC, both embedding and compulsory attachment. This was introduced in [4] in 2019 and expanded upon in a journal article entitled Formalisation and classification of grammar and template-mediated techniques to model and ontology verbalisation [5] that was published in IJMSO, and even more detail can be found in Zola Mahlaza’s recently completed PhD thesis [6]. These papers have various examples, illustrations how to categorise a system, and why one system was categorised in one way and not another. Here’s a table with several systems that combine templates and computational grammar rules and how they are categorised:
We needed a short-hand name to refer to the cumbersome and wordy description of ‘combining templates with grammar rules in a [theoretical or implemented] system in some way’, which ended up to be grammar-infused templates.
Why write about this now? Besides certain pandemic-induced priorities in 2021, the recently proposed template language for Abstract Wikipedia that I blogged about before may mix Compulsory or Partial attachment, but ought not to permit the messy embedding of grammar in a template. This may not have been clear in v1 of the proposal, but hopefully it is a little bit more so in this new version that was put online over the past few days. To make that long story short: besides a few notes at the start of its Section 3, there’s a generic description of an idea for a realization algorithm. Its details don’t matter if you don’t intend to design a new realiser from scratch and maybe not either if you want to link it to your existing system. The key take-away from that section is that there’s where the real grammar and phonological conditioning stuff happens if it’s needed. For example, for the ‘age in years’ sub-template for isiZulu, recall that’s:
Year_zu(years):"{root:Lexeme(L686326)} {concord:RelativeConcord()}{Copula()}{concord_1<nummod:NounPrefix()}-{nummod:Cardinal(years)}"
The template language sets some boundaries for declaring such a template, but it is a realiser that has to interpret ‘keywords’, such as root, concord, and RelativeConcord, and do something with it so that the output ends up correctly; in this case, from ‘year’ + ‘25’ as input data to iminyaka engama-25 as outputted text. That process might be done in line with Ariel Gutman’s realiser pipeline for Abstract Wikipedia and his proof-of-concept implementation with Scribunto or any other realizer architecture or system, such as Grammatical Framework, SimpleNLG, Ninai–Udiron, or Zola’s Nguni Grammar Engine, among several options for multilingual text generation. It might sound silly to put templates on top of the heavy machinery of a grammar engine, but it will make it more accessible to the general public so that they can specify how sentences should be generated. And, hopefully, permit a rules-as-you-go approach as well.
It is then the realiser (including grammar) engine and the partially or compulsorily attached computational grammar rules and other algorithms that work with the template. For the example, when it sees root and that the lemma fetched is a noun (L686326 is unyaka ‘year’), it also fetches the value of the noun class (a grammatical feature stored with the noun), which we always need somewhere for isiZulu NLG. It then needs to figure out to make a plural out of ‘year’, which it will know that it must do thanks to the years fetched for the instance (i.e., 25, which is plural) and the nummod that links to the root by virtue of the design and the assumption there’s a (dependency) grammar. Then, with concord:RelativeConcord, it will fetch the relative concord for that noun class, since concord also links to root. We already can do the concordial agreements and pluralising of nouns (and much more!) for isiZulu since several years. The only hurdle is that that code would need to become interoperable with the template language specification, in that our realisers will have to be able to recognise and process properly those ‘keywords’. Those words are part of an extensible set of words inspired by dependency grammars.
How this is supposed to interact smoothly is to be figured out still. Part of that is touched upon in the section about instrumentalising the template language: you could, for instance, specify it as functions in Wikifunctions that is instantly editable, facilitating an add-rules-as-you-go approach. Or it can be done less flexibly, by mapping or transforming it to another template language or to the specification of an external realiser (since it’s the principle of attachment, not embedding, of computational grammar rules).
In closing, whether the term “grammar-infused templates” will stick remains to be seen, but combining templates with grammars in some way for NLG will have a solid future at least for as long as those ML/DL-based large language model systems keep hallucinating and don’t consider languages other than English, including the intended multilingual setting for Abstract Wikipedia.
References
[1] M. Jarrar, C.M. Keet, and P. Dongilli. Multilingual verbalization of ORM conceptual models and axiomatized ontologies. STARLab Technical Report, Vrije Universiteit Brussels, Belgium. February 2006.
[2] Keet, C.M., Khumalo, L. Toward a knowledge-to-text controlled natural language of isiZulu. Language Resources and Evaluation, 2017, 51:131-157. (accepted version free access)
[3] Keet, C.M. Xakaza, M., Khumalo, L. Verbalising OWL ontologies in isiZulu with Python. The Semantic Web: ESWC 2017 Satellite Events, Blomqvist, E et al. (eds.). Springer LNCS vol 10577, 59-64. Portoroz, Slovenia, May 28 – June 2, 2017.
[4] Mahlaza, Z., Keet, C.M. A classification of grammar-infused templates for ontology and model verbalisation. 13th Metadata and Semantics Research Conference (MTSR’19). E. Garoufallou et al. (Eds.). Springer vol. CCIS 1057, 64-76. 28-31 Oct 2019, Rome, Italy.
[5] Mahlaza, Z., Keet, C.M. Formalisation and classification of grammar and template-mediated techniques to model and ontology verbalisation. International Journal of Metadata, Semantics and Ontologies, 2020, 14(3): 249-262.
[6] Mahlaza, Z. Foundations for reusable and maintainable surface realisers for isiXhosa and isiZulu. PhD Thesis, Department of Computer Science, University of Cape Town, South Africa. 2022.